Drug-Drug Interaction Analysis & Prediction
Can one predict interaction of two newly developed drugs?
Testing drugs on their compatibility is one of the major issues in nowadays pharmacology, as these interactions can significantly impact drug efficacy and patient safety. By leveraging data from DrugBank on examples of synergetic drug-drug interactions we came up with several models on how to predict newly developed drugs' interactions. This research combines natural language processing, clustering, and machine learning to tackle the complexities of DDI analysis. By embedding interaction descriptions using BioBERT, we created rich, context-aware representations for clustering interactions based on severity. Furthermore we look at possible interesting finding related to severity of interaction in "likable" drugs. Finally by encoding SMILES of each drug, we use Neural-Network models to predict a potential severity of synergetic interactions. This comprehensive approach aims to enhance our understanding of DDIs and contribute to safer, more effective pharmacological practices.
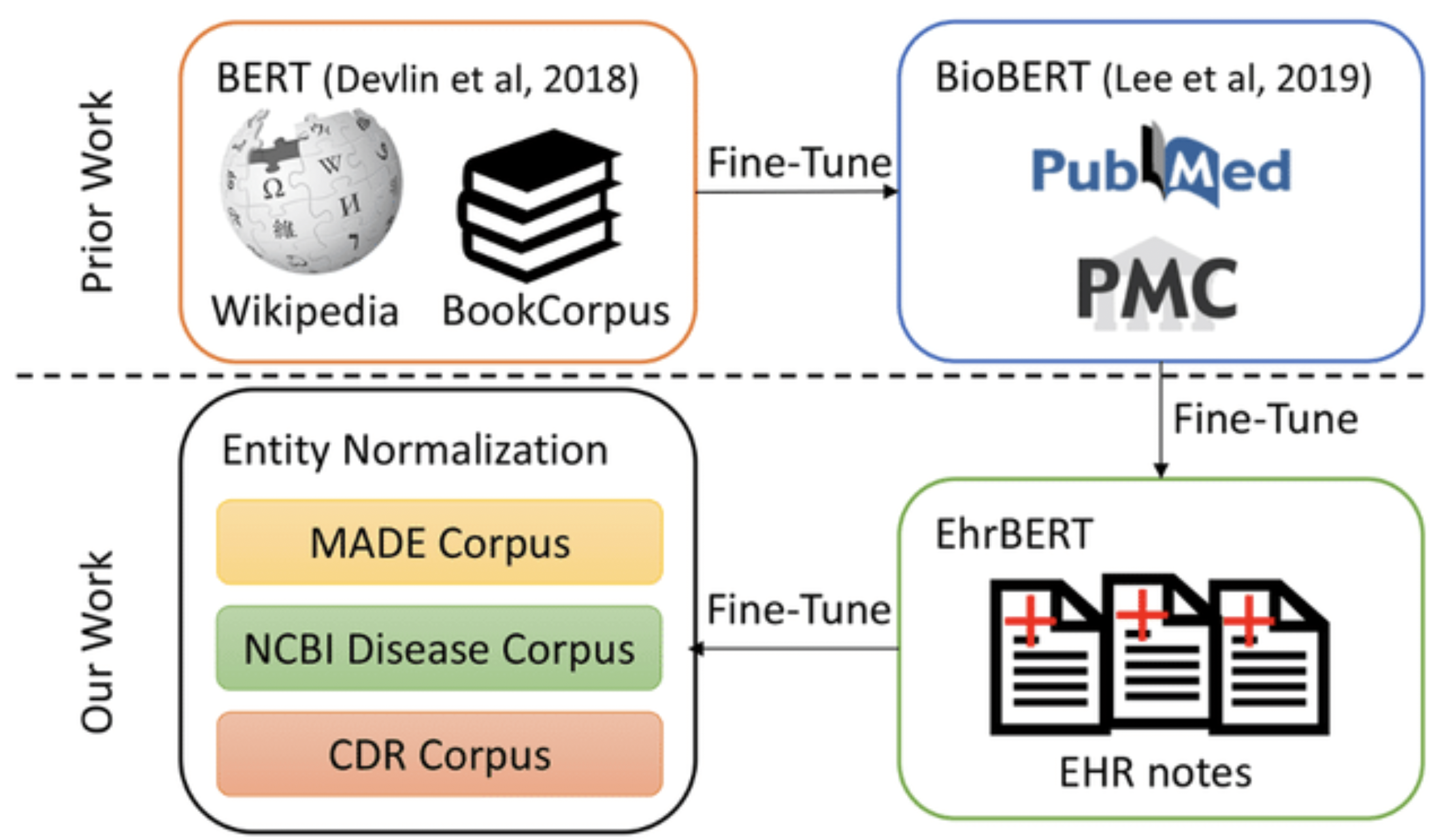
BioBert & Clustering Interactions
BioBERT (Bidirectional Encoder Representations from Transformers for Biomedical Text Mining) is a domain-specific extension of the BERT language model, designed specifically for biomedical and clinical text. Unlike standard BERT, which is trained on general language data, BioBERT incorporates large-scale biomedical corpora such as PubMed and PMC articles.
Once we utilized BioBERT to generate embeddings for each drug-drug interaction (DDI) description, we applied the K-Means algorithm to cluster the interactions based on the severity of their effects. The interactions were grouped into three main categories: Minor, Moderate, and Major. Notably, a greater number of DDIs were classified as Moderate, reflecting a prevalent level of intermediate severity in the dataset.
To further understand the characteristics of the clustered interactions, we analyzed the text descriptions within each severity category. Word clouds were generated to visualize the most frequent terms associated with ‘Minor,’ ‘Moderate,’ and ‘Major’ interactions, offering a quick insight into the dominant themes and keywords in each group.

Fingerprint Transformation using RDKit
Our prediction model is based on SMILES structure of two drugs, thus we first need to encode those molecular structures. For this purpose we used Morgan fingerprints transformation provided by RDKit that converts SMILES into 1024-bits vector encoding. These fingerprints allow to encode the presence or absence of specific molecular substructures or patterns that might be crusial in the prediction of their interaction. By applying this numerical transformations it allows molecules to be efficiently compared or used in predictive models.
Prediction Model
By taking Morgan fingerprints of two drugs as the input, we are capable of developing a model that could predict the interaction between drug pairs. We use Neural-Network approach to solve this problem. First the NN model treats each drug separately but after two sequential layers, it merges the two inputs in order to make a final prediction. Our model achieves a 97.48% overall accuracy in its prediction.
DDI of "Likable" Drugs
Our initial hypothesis suggested that “likable” drugs would have less severe interactions with other drugs. To test this, we examined the severity of interactions where at least one drug was considered “likable.” However, the results indicate that the likability of a drug does not significantly impact the severity of its interactions with other drugs. As a result, we can reject our hypothesis. Based on these findings, we believe that the most effective model for prediction is one that focuses solely on the molecular structures of each drug.
Conclusion
In this study, we explored various approaches to predict the interactions between newly developed drugs using data from DrugBank. We utilized BioBERT for embedding drug-drug interaction (DDI) descriptions, followed by clustering based on interaction severity, revealing that moderate interactions were most prevalent. Additionally, we employed SMILES-based molecular structures encoded as Morgan fingerprints and used a Neural Network model to predict the severity of drug interactions, achieving a high accuracy of 97.48%. Our hypothesis that “likable” drugs would have less severe interactions was not supported by the data, leading us to conclude that molecular structure-based models are more effective. This research provides valuable insights into DDI prediction and highlights the potential of molecular structure analysis for more accurate predictions.
